Introduction
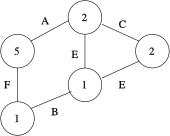
A connected, undirected and labeled graph.
The gboost toolbox is a framework for classification of connected, undirected, labeled graphs. A typical graph is shown on the right: each node and edge is labeled by a discrete value.
The gboost classifiers check for the presence of certain subgraphs in the larger graph. The subgraphs being checked are optimally determined by discriminative subgraph mining. The classification hypotheses is interpretable because only a small number of subgraphs are used to determine the overall classification decision.
On this page, we provide instructions and source code in order to foster the adoption of the graph boosting methodology. Graph Boosting has been used successfully in chemical compound classification, natural language processing and computer vision. Please see the publications section for published papers.
Features and Demo
The gboost toolbox includes code for the following functionalities:
- Discriminative Subgraph Mining
- Frequent Subgraph Mining (gSpan), code by Taku Kudo
- Subgraph-Graph isomorphism test (through VFlib)
- nu-LPBoost 2-class classifier
- nu-LPBoost 1.5-class classifier
- simple wrappers to easily train a classifier for graphs
Most of the code is written in C++ with MEX Matlab wrappers.
The following screen capture shows the output when the included example.m file from the distribution is run.
>> example
Loading example graphs...
96 training samples
69 test samples
We train a 2-class graph boosting classifier.
the settings are:
nu = 0.2, the LPBoost nu-parameter controlling training accuracy
conv_epsilon = 0.05, the LPBoost convergence tolerance parameter
max_col = 25, generate 25 hypotheses in each iteration (multiple pricing)
Please press return to start training...
=== GLPBOOST = BEGIN = 2-class graph based LPBoosting ===
LPBoost iter 1
Starting gspan-boost run...
(...)
gSpan returned 25 subgraphs
optimality: 0.34279 <= 0 + 0.05 ?
(...)
gSpan returned 25 subgraphs
optimality: 0.097308 <= 0.049357 + 0.05 ?
LPBoost optimality reached after 10 iterations.
(no improvements left)
=== END ===
The classifier has been trained successfully.
There are 161 active subgraph stumps.
Please press return to test the classifier...
test accuracy = 0.75362
test ROC AUC = 0.79899
test ROC EER = 0.75676
>>
Download
The gboost toolkit for Matlab can be downloaded below. The package includes the source code, pre-compiled binaries for the Linux/x86 and the Linux/x86-64 architectures. Also included is a set of graphs from the CPDB chemical database to test the graph boosting functionality.
Distribution: source code, precompiled binaries and demo file
- gboost-0.1.1.tar.gz (1.1Mb)
License: The software is dual-licensed under both the GNU General Public License, version 2 and the Mozilla Public License, version 1.1. A copy of the license documents is included in the distribution. The dual-licensing allows you to pick any of the two licenses and thus makes it easier to combine this software with other toolboxes.
Installation: the distribution includes a Makefile.options file which specifies the MATLABROOT variable. After editing the file accordingly, the program should compile on any recent Linux system. If you have the frequently encountered problem complaining about GCC_3.3 not being found when you run the mex functions, please refer to this discussion at the Mathworks forums.
Publications
- Weighted Substructure Mining for Image Analysis, CVPR 2007, Sebastian Nowozin, Koji Tsuda, Takeaki Uno, Taku Kudo and Gökhan BakIr.
- A Linear Programming Approach for Molecular QSAR analysis, MLG 2006, Hiroto Saigo, Tadashi Kadowaki and Koji Tsuda.
- An Application of Boosting to Graph Classification, NIPS 2004, Taku Kudo, Eisaku Maeda and Yuji Matsumoto.
- Clustering Graphs by Weighted Substructure Mining, ICML 2006, Koji Tsuda and Taku Kudo.
Contact
- nowozin@gmail.com, primary author of the Matlab toolkit
- koji.tsuda@tuebingen.mpg.de
If you have comments or questions, please feel free to contact me. Thanks!